SPARC, led by Dr. Shahid Mukhtar at Clemson University, serves as a Certified Service Provider (CSP) for Complete Genomics. Through this partnership, SPARC provides end-to-end spatial transcriptomics services using the Stereo-seq (SpaTial Enhanced REsolution Omics sequencing) platform developed by STOmics, a spatial biology division of Complete Genomics.
Why spatial, why now
Spatial omics captures gene expression in tissue context, enabling questions about cellular neighborhoods, microenvironments, and tissue architecture that cannot be answered with dissociation-only methods. Spatial technologies bridge morphology with molecular readouts, supported by dedicated workflows and analysis support.



Why Stereo-seq
Stereo-seq is a next-generation spatial transcriptomics technology that enables genome-wide mapping of gene expression directly within intact tissue sections. By capturing mRNA on densely patterned arrays of spatially barcoded DNA nanoballs, Stereo-seq preserves the physical context of gene activity while achieving extremely high spatial resolution across large tissue areas.
-
Ultra-high spatial resolution
-
Whole-transcriptome coverage
-
Large tissue field of view
-
Powerful biological insight
What SPARC offers
-
Experimental design consultation
-
Guidance on experimental design and project planning
-
Recommendations for tissue handling and sample preparation
-
Feasibility assessment for proposed studies
-
-
Spatial transcriptomics at high spatial resolution
-
Tissue intake, quality assessment, and sample handling support
-
Cryo-embedding, cryosectioning, and slide preparation
-
Staining, histology support, and imaging
-
Spatial transcriptomics workflow execution, including permeabilization and Stereo-seq
-
Spatial library preparation and sequencing coordination
-
-
Single-cell omics analysis
-
Processing and analysis of single-cell RNA-seq and related omics datasets
-
Cell clustering, cell-type identification, and annotation
-
Integration of single-cell datasets with spatial transcriptomics data
-
Generation of interpretable visualizations and reports for biological insight
-
-
Computational analysis
-
Generation of expression matrices and spatial gene expression maps
-
Clustering and spatial domain identification
-
Cell-type annotation and spatial mapping
-
Standard visualizations and summary reports using established analysis workflows
-
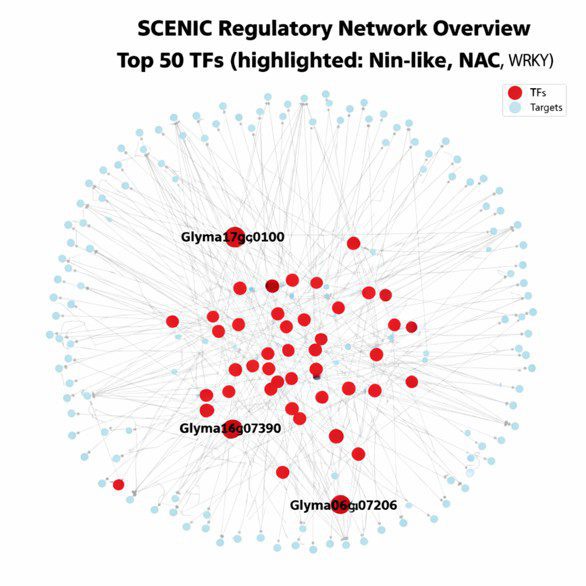
Advanced gene regulatory network (GRN analysis
-